In computer science it is common to use randomized algorithms. The same is true in statistics: there are many ways that adding randomness can make things easier. But the way that randomness enters, varies quite a bit in different methods. I thought it might be interesting to collect some specific examples of statistical procedures where added randomness plays some role. (I am not referring to the randomness inherent in the original data but, rather, I refer to randomness in the statistical method itself.)
(1) Randomization in causal inference. The mean difference between a treated group and untreated group is not, in general, equal to the causal effect
. (Correlation is not causation.) Moreover,
is not identifiable. But if we randomly assign people to the two groups then, magically,
. This is easily proved using either the directed graph approach to causation or the counterfactual approach: see here for example. This fact is so elementary that we tend to forget how amazing it is. Of course, this is the reason we spend millions of dollars doing randomized trials.
(As an aside, some people say that there is no role for randomization in Bayesian inference. In other words, the randomization mechanism plays no role in Bayes’ theorem. But this is not really true. Without randomization, we can indeed derive a posterior for but it is highly sensitive to the prior. This is just a restatement of the non-identifiability of
. With randomization, the posterior is much less sensitive to the prior. And I think most practical Bayesians would consider it valuable to increase the robustness of the posterior.)
(2) Permutation Tests. If and
and you want to test
versus
, you can get an exact, distribution-free test by using the permutation method. See here. We rarely search over all permutations. Instead, we randomly select a large number of permutations. The result is still exact (i.e. the p-value is sub-uniform under
.)
(3) The Bootstrap. I discussed the bootstrap here. Basically, to compute a confidence interval, we approximate the distribution
with the conditional distribution
where ,
and
is a sample from the empirical distribution. But the distribution
is intractable. Instead, we approximate it by repeatedly sampling from the empirical distribution function. This makes otherwise intractable confidence intervals trivial to compute.
(4) -means++. Minimizing the objective function in
-means clustering is NP-hard. Remarkably, as I discussed here, the
-means++ algorithm uses a careful randomization method for choosing starting values and gets close to the minimum with high probability.
(5) Cross-Validation. Some forms of cross-validation involve splitting the data randomly into two or more groups. We use one part(s) for fitting and the other(s) for testing. Some people seem bothered by the randomness this introduces. But it makes risk estimation easy and accurate.
(6) MCMC. An obvious and common use of randomness is random sampling from a posterior distribution, usually by way of Markov Chain Monte Carlo. This can dramatically simplify Bayesian inference.
These are the first few things that came to my mind. Are there others I should add to the list?
20 Comments
In the linked post on causal inference, you mention a future post on Simpson’s paradox and the case in which p(y|Set X = x) \neq p(y|x). The huddled masses (or maybe it’s just me) eagerly await this post!
Thanks for reminding about that.
I will post on it soon
Maybe ABC should be mentioned as a multi-level randomisation: the data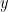 is replaced with a randomised version
is replaced with a randomised version  in the Bayes conditioning and the Bayesian inference only remains exact if the data is indeed replaced with its (once) randomised version. This comes in addition to the Monte Carlo randomisation that underlines your examples (2), (3), (4), and (6). Even though I would have argued at the classification of those as randomisation methods, per se, had we had the luck to meet in Roma last week…!
in the Bayes conditioning and the Bayesian inference only remains exact if the data is indeed replaced with its (once) randomised version. This comes in addition to the Monte Carlo randomisation that underlines your examples (2), (3), (4), and (6). Even though I would have argued at the classification of those as randomisation methods, per se, had we had the luck to meet in Roma last week…!
I knew I would regret missing our meeting in Rome!
Btw, I loved your biosketch in amstat news: “In his spare time, he enjoys mountain climbing, parachuting, and big game hunting.”
Yes I was able to sneak that by
Randomness can also be an effective regularizer during training. Random Forest randomly decides to ignore a large subset of features at every split decision. Geoffrey Hinton’s dropout technique for regularization in training neural networks does something similar.
Multiple Imputation.
Randomly shuffling the examples can speed-up optimization in the finite sample case. Here are a couple of papers mentioning this (as a disclaimer, I’m an author of one of them):
http://arxiv.org/abs/1209.1873
http://arxiv.org/abs/1202.6258
There was at least another one I knew of but I can’t seem to find it. I’m pretty sure it was by Léon Bottou.
This is an attention-catching, but misleading, title! An appropriate title would have been “The Value of Using Randomization”
There’s a few further techniques that crossed my mind, including optimization algorithms such as SA, GA (I assume if you count MCMC you should count them as well) as well as various re-sampling or simulation-based null-models.
Maybe useful to classify in 1) Monte-Carlo algorithms, including MCMC, SA, GA and heaps of other ML algorithms 2) Algorithms that re-sample or permute data, including bootstrapping and permutation null models 3) Stochastic simulations, including simulation-based null models, ABC, etc. , ? Not sure whether that covers everything …
Excellent post. (I take it (1) would be qualified for “the” causal effect in the population randomly sampled.) But my main point, rather than adding examples, is to draw attention to the deep implications of these. Notably, they immediately scotch the oft repeated charge that sampling distributions and associated frequentist methods are valuable only in some long run: it is precisely the counterfactual reasoning that enables them to be relevant to the process, or data generating source, at hand. A causal question, say, entails counterfactual claims about “what it would be like were…” and the sampling distribution lets us infer “what it would be like were…” using the sample. Therein lies the key that opens the door to ampliative inference (whichever school takes advantage of them)—or so I have been claiming.
I recently came across the “Pushed” confidence intervals of Lorden (as described in a PhD thesis by T. V. Asparouhov). These are randomized confidence intervals on the Bernoulli parameter. You take the number of successes observed in the experiment, add a uniform random variate in [-0.5,+0.5] and then use that value to construct the confidence interval. Asparouhov claims these intervals are more efficient than non-randomized ones.
Speaking of randomized algorithms, a big theoretical question in computer science is whether randomization plays any ESSENTIAL role in allowing some things to be computed in polynomial time. There is a whole field of “de-randomization”, which seeks to convert randomized algorithms into deterministic ones. There are some formal results where randomness helps, but there are many open questions!
A simple example: jittering.
I’d appreciate some clarification of your assertion: “This [the effectiveness of randomisation for causal inference] is easily proved using either the directed graph approach to causation or the counterfactual approach”. However, approaches such as these can do no more than provide a system for inferring conclusions from assumptions, either explicit or implicit. Exactly what assumptions are required to infer this specific conclusion?
I’m not sure what you are asking Phil.
Randomization turns a non-identifiable parameter
into a identifiable parameter.
But I said this in the post so perhaps I am not understanding your question.
Random Forests!
And Meinshausen’s stability subsampling method for model selection.
In
“C. Hennig and T. F. Liao How to find an appropriate clustering for mixed type variables with application to socioeconomic stratification. Journal of the Royal Statistical Science, Series C (Applied Statistics) 62, 309-369 (2013), with discussion”
we observed that maximising certain cluster validation indexes (average silhouette width, Calinski/Harabasz; the phenomenon may apply to others) as recommeded in the literature led to an estimate of the number of clusters for which the clustering is not significantly better than random data simulated from a null model for “no clustering but some other realistic structure”. However, other supposedly non-optimal numbers of clusters led to significantly better clusterings.
Comparing the validation index values of clusterings to what happens under such null models is generally helpful for cluster validation.
Generally it may be worthwhile when interested in some kind of structure to compare with a null model that incorporates other kinds of structure that one could legitimately expect in such data, instead testing against an unrealistically simplistic null model such as iid normal.
Obviously, though, adding randomness here is only required because I am (“we all are?”) too stupid to compute such things from sufficiently flexible null models theoretically. (But this applies to some other examples listed here, too.)
Randomized response — that old trick for getting people in aggregate to reveal their preferences for sensitive questions.
One Trackback
[…] Larry Wasserman in 2013: […]